SAS Thread - esasspec - XMM-Newton
Creation of EPIC ESAS spectral analysis files for a cluster radial profile
|
Introduction This thread describes how to create EPIC spectral and response files for a complicated multi-region analysis combining data from the three instruments. This thread uses data from the observation of Abell 1795, ObsID 0097820101, which is also used as the Spectral Analysis example where a script and output files are provided. Expected Outcome The final outcome of this thread is the spectral analysis of the radial profile of the cluster of galaxies Abell 1795 with ten annuli. SAS Tasks to be Used
Prerequisites Useful Links
Last Reviewed: 31 January 2025, for SAS v22.0Last Updated: 25 May 2023 |
Procedure
This thread contains a step-by-step recipe to create EPIC background and source spectra and response files from all three instruments.
-
Set up your SAS environment (following the SAS Startup Thread)
-
Create MOS and pn (including OOT processing) event files for your observation using emchain and epchain:
epchain
epchain withoutoftime=true
emchainNote that epchain, by default, only does the first pn segment that it finds. Sometimes this is not all the pn data that there is. To determine how many pn data segments there are:
epchain exposure=99
Since there probably aren't 99 exposures, this will produce an error statement. It will also produce a list of pn segments:
-:--------------------------------------------------------------------
-:- ARGV = ! exposure=99 !
-:- Test: TIME RAWX RAWY DETX DETY X Y PHA PI FLAG PATTERN PAT_ID PAT_SEQ OFFSETX
-:- Tested: TIME RAWX RAWY DETX DETY X Y PHA PI FLAG PATTERN PAT_ID PAT_SEQ OFFSETX
-:- Test: double int16 int16 int16 int16 int32 int32 int16 int16 int32 int8 int16 int8 int16
-:- Tested: double int16 int16 int16 int16 int32 int32 int16 int16 int32 int8 int16 int8 int16
-:- EXPOSURE = 99
-:- 2 IMAGING exposures in ODF:
-:- 1: 0100_0097820101_PNS005 IM
-:- 2: 0100_0097820101_PNS013 IM
** -: error (exposure), exposure not found in /Users/kuntz/Data/xmm_nacho/0097820101/ODF directory. Check exposure index 99You can process the nth segment with:
epchain exposure=n
-
Determine which MOS chips are in anomalous states using emanom:
emanom eventfile=mos1S003.fits
emanom eventfile=mos2S004.fitsNote that emanom will indicate which CCDs are operating in an anomalous mode (B) or are not operating at all (O). These CCDs should be excluded in downstream processing. Whether or not a CCD is operating in an anomalous mode is not always clear (I or U). For those CCDs the user should consider the particular impact on their own science when deciding to include or exclude the chip for further processing.
1S003
M1 CCD: 2 Hard = 4.64406776 +/- 0.666529477 Status: G
M1 CCD: 3 Hard = 5.26829290 +/- 0.897465289 Status: G
M1 CCD: 4 Hard = 2.76271176 +/- 0.296809137 Status: I
M1 CCD: 5 Hard = 1.59555554 +/- 0.135668769 Status: B
M1 CCD: 6 Hard = 4.12727261 +/- 0.620288074 Status: G
M1 CCD: 7 Hard = 4.73684216 +/- 0.690467894 Status: G
Status: G is good at all energies
: I is intermediate for E<1 keV
: B is bad for E<1 keV
: O is off, chip not in use
: U is undetermined (low band counts ≤ 0)2S004
M2 CCD: 2 Hard = 2.83838391 +/- 0.331735671 Status: G
M2 CCD: 3 Hard = 3.95454550 +/- 0.544851601 Status: G
M2 CCD: 4 Hard = 7.06976748 +/- 1.15185773 Status: G
M2 CCD: 5 Hard = 3.69565225 +/- 0.434309751 Status: I
M2 CCD: 6 Hard = 3.96721315 +/- 0.568374038 Status: G
M2 CCD: 7 Hard = 3.88157892 +/- 0.499318421 Status: G
Status: G is good at all energies
: I is intermediate for E<1 keV
: B is bad for E<1 keV
: O is off, chip not in use
: U is undetermined (low band counts ≤ 0) -
Determine which time intervals are contaminated by soft proton flares and remove them to create cleaned event files for your observation with espfilt:
espfilt eventfile=mos1S003.fits method=histogram withsmoothing=yes smooth=51 rangescale=6.0 \
allowsigma=3.0
espfilt eventfile=mos2S004.fits method=histogram withsmoothing=yes smooth=51 rangescale=6.0 \
allowsigma=3.0
espfilt eventfile=pnS003.fits method=histogram withsmoothing=yes smooth=51 rangescale=15.0 \
allowsigma=3 withoot=Y ootfile=${PN}-oot.fitsespfilt filters an event file to remove periods with high soft proton flux. This routine is not fool-proof, and may fail if too much of the observation contains soft proton flares. Note that the call to espfilt for the pn is slightly different than the call for the MOS. It is essential to examine the diagnostic plots created by espfilt as they provide an indication of the quality of the data. These files have names like mos1S003-hist.qdp, mos2S004-hist.qdp, and pnS005-hist.qdp and can be plotted using the command, for example, qdp mos1S003-hist.qdp. Note that while much of the soft proton contamination is removed, there is likely to be some residual contamination left. Figure 1 shows the light-curve screening for the pn instrument.
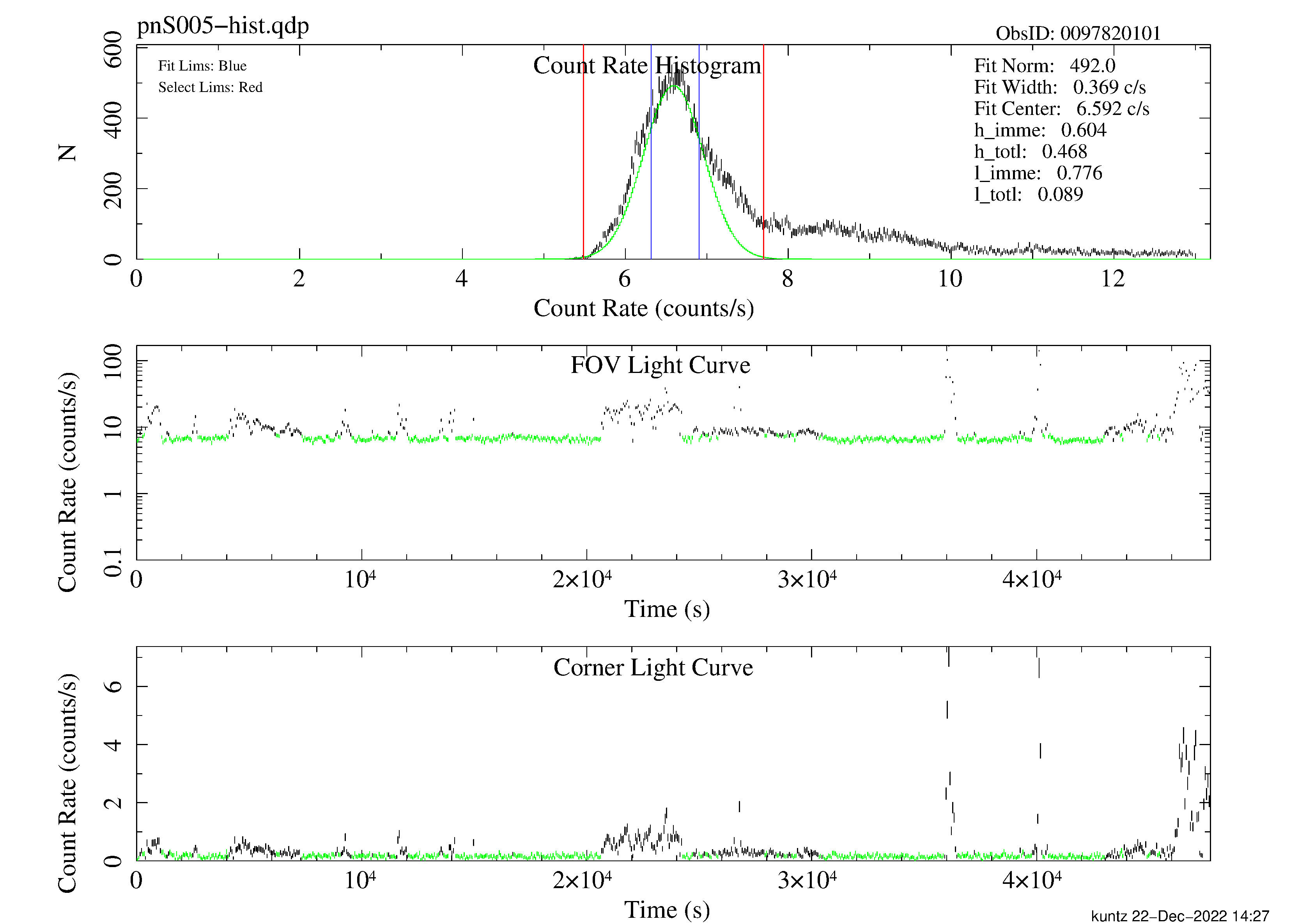
Figure 1: pn light curve for the Abell 1795 observation. Notice the residual variation of the nominally good data which indicates the possible existence of residual soft proton contamination. Note also that the pn detector is more sensitive to the soft protons than are the MOS detectors.
-
Run source detection and make point-source masks with cheese
cheese mos1file=mos1S003-allevc.fits mos2file=mos2S004-allevc.fits pnfile=pnS003-allevc.fits \
pnootfile=pnS003-allevcoot.fits elowlist=400 ehighlist=7200 ratetotal=1.0 dist=50. \
keepinterfiles=false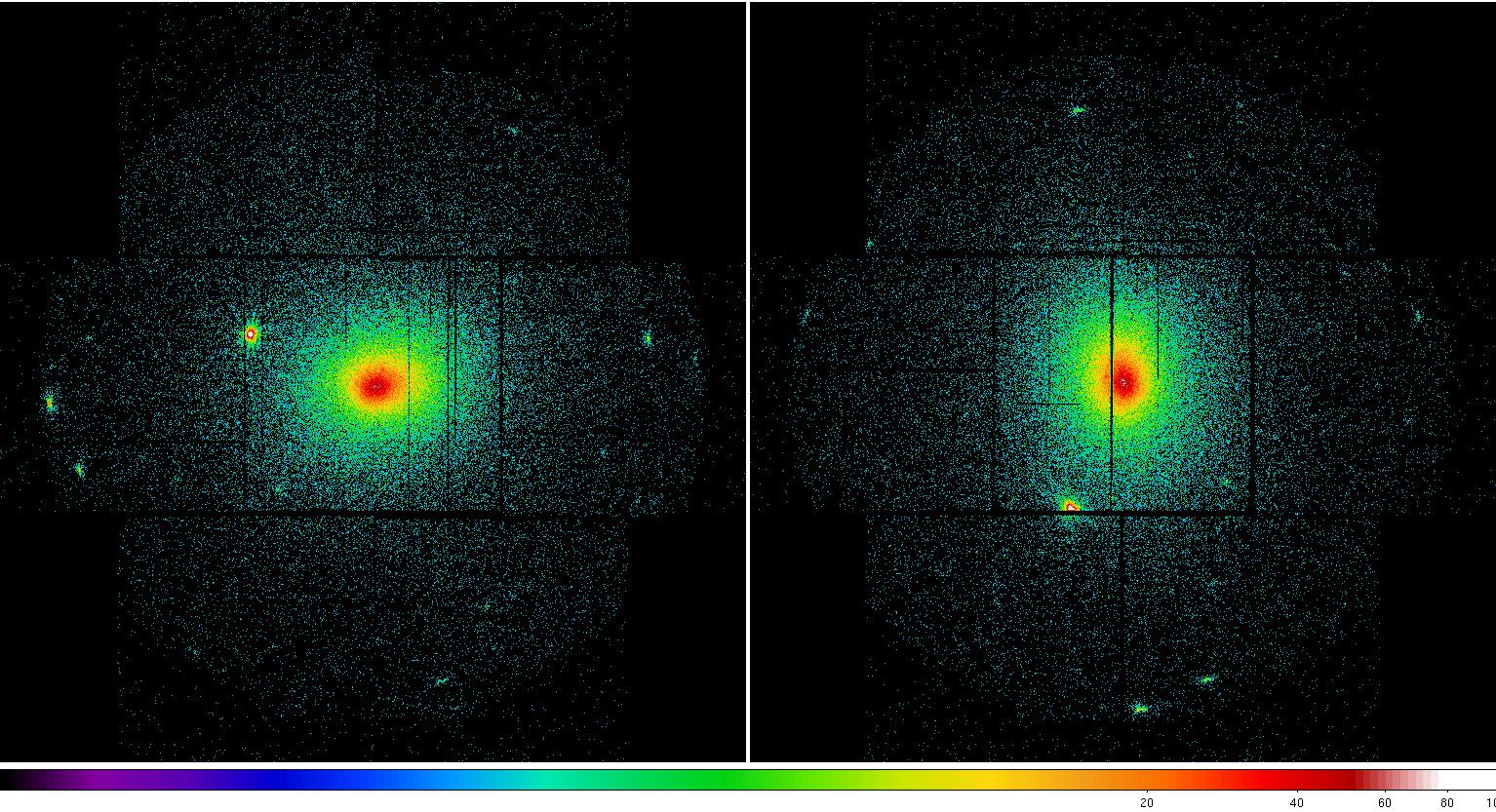
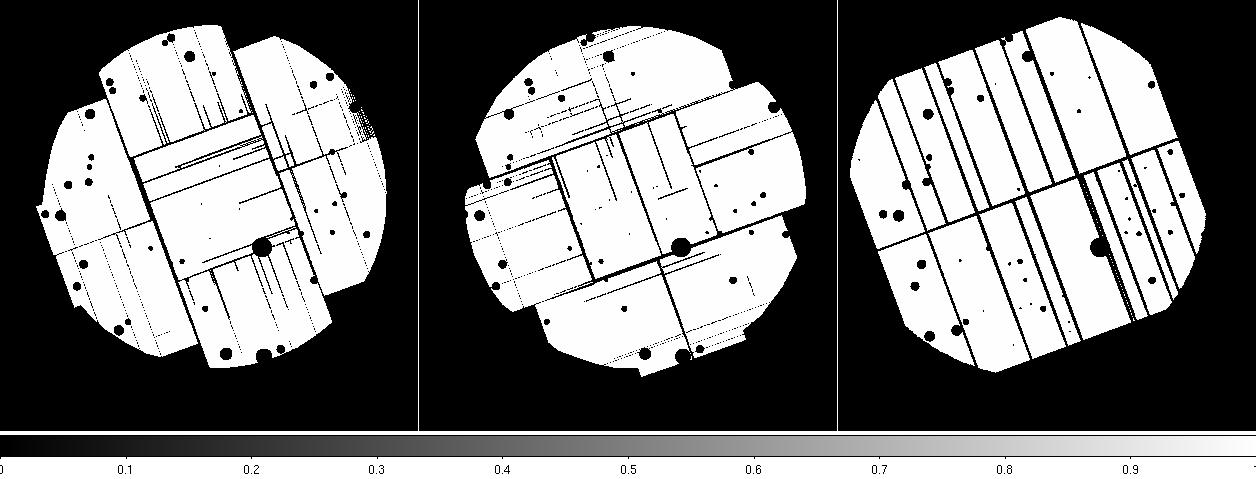
Figure 2: MOS1 (left) and MOS2 (right) images in the soft (0.2-1.0 keV) band. Note the excess counts in the upper left CCD in the MOS1 image most noticeable in the unexposed (to the sky) upper left corner. Displayed by the command: ds9 *soft* &.
-
Use mosspectra and pnspectra to create the required intermediate spectra (for the entire region of interest as well as spectra from the individual CCDs), RMF and ARF files, and detector images for one band and three detectors. MOS CCDs affected by anomalous states should be deselected (this is done by the ccds parameter). For example, MOS1 CCD6 should be deselected if the observation took place after the meteorite damage. The region selection expression is input through the withregion and regionfile parameters. The region selection expression should be in detector coordinates. If withregion=false or the input file (regmos1.txt in this case) does not exist, the default is to process the entire FOV. The input energies are in eV.
mosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=regmos1.txt pattern=12 \
withsrcrem=yes maskdet=mos1S003-bkgregtdet.fits masksky=mos1S003-bkgregtsky.fits \
elow=400 ehigh=7200 ccds="T T T F F T T"
mosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=regmos2.txt pattern=12 \
withsrcrem=yes maskdet=mos2S004-bkgregtdet.fits masksky=mos2S004-bkgregtsky.fits \
elow=400 ehigh=7200 ccds="T T T T F T T"
pnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits withregion=yes \
regionfile="regpn.txt" pattern=0 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits \
masksky=pnS005-bkgregtsky.fits elow=400 ehigh=7200 quads="T T T T"Note: successive runs of mosspectra or pnspectra may overwrite previously constructed files. From this point on, one can run the reduction for several different energy bands in parallel. However, changing regions would require starting over from this point. Thus, it is a good idea to keep note of what files exist before mosspectra so you can return to this point of the reduction if you need to.
-
Use mosback and pnback to create the quiescent particle background (QPB) spectra and images (in detector coordinates). mosback and pnback create QDP plot files which shows the source and model background spectra for the observation. Any discrepancies at higher energies probably indicate residual soft proton contamination, unless there are really hard and bright sources in the field. In the case of this observation the discrepancy at high energies is consistent with soft protons as a residual contamination was already expected from the light curve histogram. The QDP files have names like mos1S005-spec.qdp. The same CCD selection must be used here as were used in mosspectra and pnspectra.
mosback inspecfile=mos1S003-fovt.pi elow=400 ehigh=7200 ccds="T T T T T T T"
mosback inspecfile=mos2S004-fovt.pi elow=400 ehigh=7200 ccds="T T T T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi \
elow=400 ehigh=7200 quads="T T T T"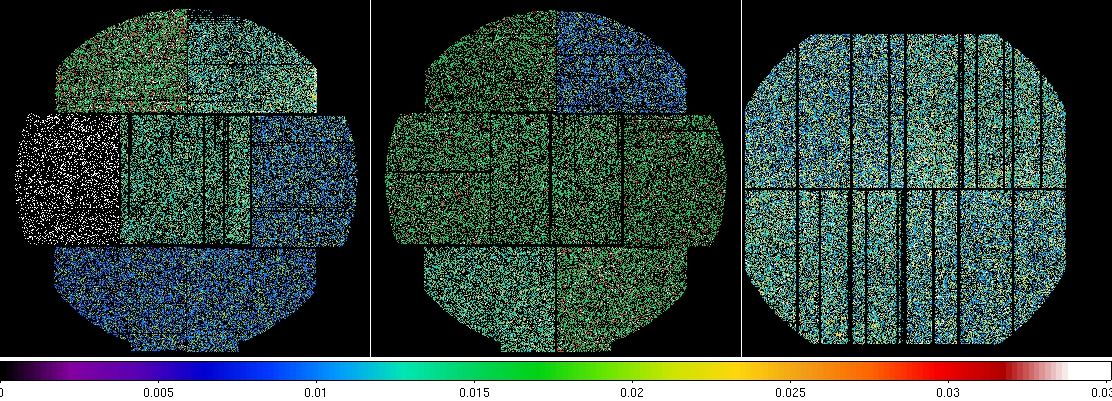
Figure 4: MOS1 (left), MOS2 (middle), and pn (right) model particle images in the soft (0.4-7.2 keV) band. The different colors for the different CCDs are an indication of the variations in their exposures. Displayed by the command: ds9 mos1S003-bkgimdet-400-7200.fits mos2S004-bkgimdet-400-7200.fits pnS005-bkgimdet-400-7200.fits &
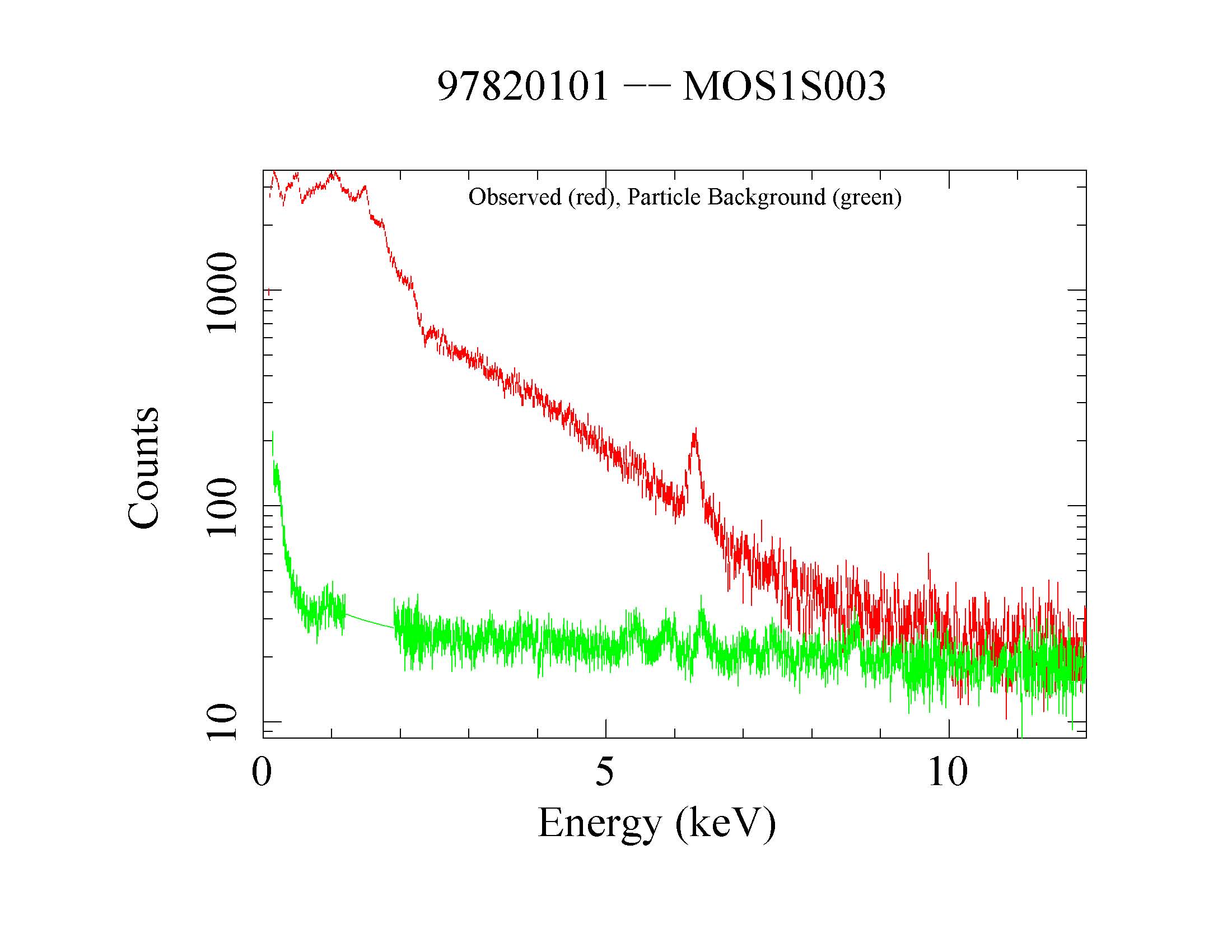
Figure 5: MOS1 source (red) and QPB (green) spectra. Displayed by the command: qdp mos1S003-bkgspec.qdp.
Note that the output images from mosback and pnback are in detector coordinates and need to be rotated into sky coordinates with rotdet2sky:
rotdet2sky intemplate=mos1S003-fovimsky-400-7200.fits inimage=mos1S003-bkgimdet-450-1250 \
outimage=mos1S003-bkgimsky-400-7200.fits withdetxy=false withskyxy=false
rotdet2sky intemplate=mos2S004-fovimsky-400-7200.fits inimage=mos2S004-bkgimdet-450-1250 \
outimage=mos2S004-bkgimsky-400-7200.fits withdetxy=false withskyxy=false
rotdet2sky intemplate=pnS005-fovimsky-400-7200.fits inimage=pnS005-bkgimdet-450-1250 \
outimage=pnS005-bkgimsky-400-7200.fits withdetxy=false withskyxy=false -
Make images of the FOV without the point sources
combimage combines the MOS1, MOS2, and pn images, as well as images from multiple exposures. Since the spectra and images were created without point sources, combimage must be run using the cheese masking. Note that the soft proton background is not removed for this image. See the ESAS Imaging thread for more information.
combimage prefixlist='1S003 2S004 S005' withpartbkg=true withspbkg=false withswcxbkg=false withcheese=true cheesetype=t \
elowlist=400 ehighlist=7200binadapt adaptively smooths the image.
bindapt prefix=comb elow=400 ehigh=7200 withpartbkg=true withspbkg=false \
withswcxbkg=false withmask=false withsmoothing=true smoothcounts=50 \
withbinning=true binfactor=2 maskthresh=0.02Save your images containing the point sources so they don't get overwritten. (In fact, it would be a good idea to save all of the products of mosspectra/pnspectra and mosback/pnback!)
mv comb-fovimsky-400-7200.fits comb-fovimsky-400-7200-nps.fits
mv comb-bkgimsky-400-7200.fits comb-bkgimsky-400-7200-nps.fits
mv comb-expimsky-400-7200.fits comb-expimsky-400-7200-nps.fits
mv comb-rateimsky-400-7200.fits comb-rateimsky-400-7200-nps.fits
mv comb-sigimsky-400-7200.fits comb-sigimsky-400-7200-nps.fits -
It is often quite often useful to create and image of the entire field of view both with and without point sources, to get a perspective on the observation. This section is a repetition of the previous step with suitable changes in parameters.
mosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=regmos1.txt \
pattern=12 withsrcrem=no elow=400 ehigh=7200 ccds="T T T T T T T"
mosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=regmos2.txt \
pattern=12 withsrcrem=no elow=400 ehigh=7200 ccds="T T T T T T T"
pnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile="regpn.txt" pattern=0 withsrcrem=no elow=400 ehigh=7200 \
quads="T T T T"mosback inspecfile=mos1S003-fovt.pi elow=400 ehigh=7200 ccds="T T T T T T T"
mosback inspecfile=mos2S004-fovt.pi elow=400 ehigh=7200 ccds="T T T T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi elow=400 ehigh=7200 \
quads="T T T T"rotdet2sky intemplate=mos1S003-fovimsky-400-7200.fits inimage=mos1S003-bkgimdet-450-1250 \
outimage=mos1S003-bkgimsky-400-7200.fits withdetxy=false withskyxy=false
rotdet2sky intemplate=mos2S004-fovimsky-400-7200.fits inimage=mos2S004-bkgimdet-450-1250 \
outimage=mos2S004-bkgimsky-400-7200.fits withdetxy=false withskyxy=false
rotdet2sky intemplate=pnS005-fovimsky-400-7200.fits inimage=pnS005-bkgimdet-450-1250 \
outimage=pnS005-bkgimsky-400-7200.fits withdetxy=false withskyxy=falsecombimage prefixlist='1S003 2S004 S005' withpartbkg=true withspbkg=false \
withswcxbkg=false withcheese=false elowlist=400 ehighlist=7200bindapt prefix=comb elow=400 ehigh=7200 withpartbkg=true withspbkg=false \
withswcxbkg=false withmask=false withsmoothing=true smoothcounts=50 \
withbinning=true binfactor=2 maskthresh=0.02mv comb-fovimsky-400-7200.fits comb-fovimsky-400-7200-ps.fits
mv comb-bkgimsky-400-7200.fits comb-bkgimsky-400-7200-ps.fits
mv comb-expimsky-400-7200.fits comb-expimsky-400-7200-ps.fits
mv comb-rateimsky-400-7200.fits comb-rateimsky-400-7200-ps.fits
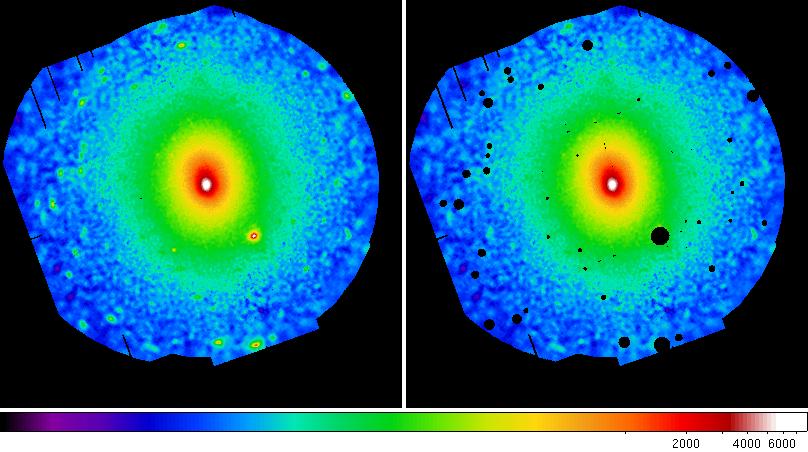
mv comb-sigimsky-400-7200.fits comb-sigimsky-400-7200-ps.fitsDisplay and compare the smoothed images.
ds9 comb-rateimsky-400-7200-ps.fits comb-rateimsky-400-7200-nps.fits &
Figure 4 shows the final adaptively smoothed, QPB subtracted, and exposure corrected images with and without point sources excluded.
-
Before we start creating the annuli, we need to clean things up and remove many of the products of mosspectra/pnspectra and mosback/pnback out of the way.
rm -f *fovimsky.fits *fovimspdet.fits *expimsky.fits *maskimsky.fits *fovt.pi \
*fovtoot.pi *rmf *arf -
Use xmmselect to find the detector coordinates of the center of the cluster for each of the three detectors. Use these coordinates to create the region selection expressions for the annuli. This is a bit tedious. For this thread the annuli are: 0-30", 30-60", 60-120", 120-180", 180-240", 240-300", 300-420", 420-540", 540-720", and 720-840". The region files used below have names starting with reg1, reg2, and reg3 for the MOS1, MOS2, and pn detectors, respectively. Note that the units are 0.05 arc seconds.
reg1-0-30.txt: &&((DETX,DETY) IN circle(201.0,-219.0,600))
reg1-30-60.txt: &&((DETX,DETY) IN circle(201.0,-219.0,1200))&& \
!((DETX,DETY) IN circle(201.0,-219.0,600))
reg1-60-120.txt: &&((DETX,DETY) IN circle(201.0,-219.0,2400))&& \
!((DETX,DETY) IN circle(201.0,-219.0,1200))
reg1-120-180.txt: &&((DETX,DETY) IN circle(201.0,-219.0,3600))&&\
!((DETX,DETY) IN circle(201.0,-219.0,2400))
reg1-180-240.txt: &&((DETX,DETY) IN circle(201.0,-219.0,4800))&& \
!((DETX,DETY) IN circle(201.0,-219.0,3600))
reg1-240-300.txt: &&((DETX,DETY) IN circle(201.0,-219.0,6000))&& \
!((DETX,DETY) IN circle(201.0,-219.0,4800))
reg1-300-420.txt: &&((DETX,DETY) IN circle(201.0,-219.0,8400))&& \
!((DETX,DETY) IN circle(201.0,-219.0,6000))
reg1-420-540.txt: &&((DETX,DETY) IN circle(201.0,-219.0,10800))&& \
!((DETX,DETY) IN circle(201.0,-219.0,8400))
reg1-540-720.txt: &&((DETX,DETY) IN circle(201.0,-219.0,14400))&& \
!((DETX,DETY) IN circle(201.0,-219.0,10800))
reg1-720-840.txt: &&((DETX,DETY) IN circle(201.0,-219.0,16800))&& \
!((DETX,DETY) IN circle(201.0,-219.0,14400))
reg2-0-30.txt: &&((DETX,DETY) IN circle(39.5,-26.0,600))
reg2-120-180.txt: &&((DETX,DETY) IN circle(39.5,-26.0,3600))&& \
!((DETX,DETY) IN circle(39.5,-26.0,2400))
reg2-180-240.txt: &&((DETX,DETY) IN circle(39.5,-26.0,4800))&& \
!((DETX,DETY) IN circle(39.5,-26.0,3600))
reg2-240-300.txt: &&((DETX,DETY) IN circle(39.5,-26.0,6000))&& \
!((DETX,DETY) IN circle(39.5,-26.0,4800))
reg2-300-420.txt: &&((DETX,DETY) IN circle(39.5,-26.0,8400))&& \
!((DETX,DETY) IN circle(39.5,-26.0,6000))
reg2-30-60.txt: &&((DETX,DETY) IN circle(39.5,-26.0,1200))&& \
!((DETX,DETY) IN circle(39.5,-26.0,600))
reg2-420-540.txt: &&((DETX,DETY) IN circle(39.5,-26.0,10800))&& \
!((DETX,DETY) IN circle(39.5,-26.0,8400))
reg2-540-720.txt: &&((DETX,DETY) IN circle(39.5,-26.0,14400))&& \
!((DETX,DETY) IN circle(39.5,-26.0,10800))
reg2-60-120.txt: &&((DETX,DETY) IN circle(39.5,-26.0,2400))&& \
!((DETX,DETY) IN circle(39.5,-26.0,1200))
reg2-720-840.txt: &&((DETX,DETY) IN circle(39.5,-26.0,16800))&& \
!((DETX,DETY) IN circle(39.5,-26.0,14400))
reg3-0-30.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,600))
reg3-120-180.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,3600))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,2400))
reg3-180-240.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,4800))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,3600))
reg3-240-300.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,6000))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,4800))
reg3-300-420.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,8400))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,6000))
reg3-30-60.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,1200))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,600))
reg3-420-540.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,10800))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,8400))
reg3-540-720.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,14400))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,10800))
reg3-60-120.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,2400))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,1200))
reg3-720-840.txt: &&((DETX,DETY) IN circle( 59.0, -147.5,16800))&& \
!((DETX,DETY) IN circle( 59.0, -147.5,14400)) -
Use mosspectra and pnspectra to create the intermediate spectra that are used by mosback and pnback to create the QPB spectra for the regions of interest and their associated RMF and ARF files. MOS CCDs affected by anomalous states should be deselected (this is done by the ccd# parameters). MOS1 CCD6 should be deselected if the observation took place after the meteorite damage (See Section 3.3 of the XMM-Newton Users Handbook). In order to avoid overwriting important files, the processing of an individual region should be done together and the files renamed. Images are formed by default, but need not be retained.
Skip to the end of the this section.
mosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-0-30.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-0-30.pi
mv mos1S003-bkg.pi mos1S003-bkg-0-30.pi
mv mos1S003.rmf mos1S003-0-30.rmf
mv mos1S003.arf mos1S003-0-30.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-0-30.fits
specgroup spectrumset=mos1S003-obj-0-30.pi mincounts=100 rmfset=mos1S003-0-30.rmf \
arfset=mos1S003-0-30.arf backgndset=mos1S003-bkg-0-30.pi \
groupedset=mos1S003-obj-0-30-grp.pimosspectra prefix=2S004 caldb=/XMM/sas/CCF/esas/ region=reg2-0-30.txt\
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-0-30.pi
mv mos2S004-bkg.pi mos2S004-bkg-0-30.pi
mv mos2S004.rmf mos2S004-0-30.rmf
mv mos2S004.arf mos2S004-0-30.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-0-30.fits
specgroup spectrumset=mos2S004-obj-0-30.pi mincounts=100 rmfset=mos2S004-0-30.rmf \
arfset=mos2S004-0-30.arf backgndset=mos2S004-bkg-0-30.pi \
groupedset=mos2S004-obj-0-30-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-0-30.txt mask=1 \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-0-30.pi
mv pnS005-fovt.pi pnS005-obj-0-30.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-0-30.pi
mv pnS005-bkg.pi pnS005-bkg-0-30.pi
mv pnS005.rmf pnS005-0-30.rmf
mv pnS005.arf pnS005-0-30.arf
mv pnS005-fovimspdet.fits pnS005-sp-0-30.fits
specgroup spectrumset=pnS005-obj-0-30.pi mincounts=100 rmfset=pnS005-0-30.rmf \
arfset=pnS005-0-30.arf backgndset=pnS005-bkg-0-30.pi \
groupedset=pnS005-obj-0-30-grp.pimosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-30-60.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-30-60.pi
mv mos1S003-bkg.pi mos1S003-bkg-30-60.pi
mv mos1S003.rmf mos1S003-30-60.rmf
mv mos1S003.arf mos1S003-30-60.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-30-60.fits
specgroup spectrumset=mos1S003-obj-30-60.pi mincounts=100 rmfset=mos1S003-30-60.rmf \
arfset=mos1S003-30-60.arf backgndset=mos1S003-bkg-30-60.pi \
groupedset=mos1S003-obj-30-60-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-30-60.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-30-60.pi
mv mos2S004-bkg.pi mos2S004-bkg-30-60.pi
mv mos2S004.rmf mos2S004-30-60.rmf
mv mos2S004.arf mos2S004-30-60.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-30-60.fits
specgroup spectrumset=mos2S004-obj-30-60.pi mincounts=100 rmfset=mos2S004-30-60.rmf \
arfset=mos2S004-30-60.arf backgndset=mos2S004-bkg-30-60.pi \
groupedset=mos2S004-obj-30-60-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-30-60.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-30-60.pi
mv pnS005-fovt.pi pnS005-obj-30-60.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-30-60.pi
mv pnS005-bkg.pi pnS005-bkg-30-60.pi
mv pnS005.rmf pnS005-30-60.rmf
mv pnS005.arf pnS005-30-60.arf
mv pnS005-fovimspdet.fits pnS005-sp-30-60.fits
specgroup spectrumset=pnS005-obj-30-60.pi mincounts=100 rmfset=pnS005-30-60.rmf \
arfset=pnS005-30-60.arf backgndset=pnS005-bkg-30-60.pi \
groupedset=pnS005-obj-30-60-grp.pimosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-60-120.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-60-120.pi
mv mos1S003-bkg.pi mos1S003-bkg-60-120.pi
mv mos1S003.rmf mos1S003-60-120.rmf
mv mos1S003.arf mos1S003-60-120.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-60-120.fits
specgroup spectrumset=mos1S003-obj-60-120.pi mincounts=100 rmfset=mos1S003-60-120.rmf \
arfset=mos1S003-60-120.arf backgndset=mos1S003-bkg-60-120.pi \
groupedset=mos1S003-obj-60-120-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-60-120.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-60-120.pi
mv mos2S004-bkg.pi mos2S004-bkg-60-120.pi
mv mos2S004.rmf mos2S004-60-120.rmf
mv mos2S004.arf mos2S004-60-120.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-60-120.fits
specgroup spectrumset=mos2S004-obj-60-120.pi mincounts=100 rmfset=mos2S004-60-120.rmf \
arfset=mos2S004-60-120.arf backgndset=mos2S004-bkg-60-120.pi \
groupedset=mos2S004-obj-60-120-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-60-120.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-60-120.pi
mv pnS005-fovt.pi pnS005-obj-60-120.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-60-120.pi
mv pnS005-bkg.pi pnS005-bkg-60-120.pi
mv pnS005.rmf pnS005-60-120.rmf
mv pnS005.arf pnS005-60-120.arf
mv pnS005-fovimspdet.fits pnS005-sp-60-120.fits
specgroup spectrumset=pnS005-obj-60-120.pi mincounts=100 rmfset=pnS005-60-120.rmf \
arfset=pnS005-60-120.arf backgndset=pnS005-bkg-60-120.pi \
groupedset=pnS005-obj-60-120-grp.piSkip to the end of the this section.
mosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-120-180.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-120-180.pi
mv mos1S003-bkg.pi mos1S003-bkg-120-180.pi
mv mos1S003.rmf mos1S003-120-180.rmf
mv mos1S003.arf mos1S003-120-180.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-120-180.fits
specgroup spectrumset=mos1S003-obj-120-180.pi mincounts=100 rmfset=mos1S003-120-180.rmf \
arfset=mos1S003-120-180.arf backgndset=mos1S003-bkg-120-180.pi \
groupedset=mos1S003-obj-120-180-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-120-180.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-120-180.pi
mv mos2S004-bkg.pi mos2S004-bkg-120-180.pi
mv mos2S004.rmf mos2S004-120-180.rmf
mv mos2S004.arf mos2S004-120-180.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-120-180.fits
specgroup spectrumset=mos2S004-obj-120-180.pi mincounts=100 rmfset=mos2S004-120-180.rmf \
arfset=mos2S004-120-180.arf backgndset=mos2S004-bkg-120-180.pi \
groupedset=mos2S004-obj-120-180-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-120-180.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-120-180.pi
mv pnS005-fovt.pi pnS005-obj-120-180.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-120-180.pi
mv pnS005-bkg.pi pnS005-bkg-120-180.pi
mv pnS005.rmf pnS005-120-180.rmf
mv pnS005.arf pnS005-120-180.arf
mv pnS005-fovimspdet.fits pnS005-sp-120-180.fits
specgroup spectrumset=pnS005-obj-120-180.pi mincounts=100 rmfset=pnS005-120-180.rmf \
arfset=pnS005-120-180.arf backgndset=pnS005-bkg-120-180.pi \
groupedset=pnS005-obj-120-180-grp.pimosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-180-240.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-180-240.pi
mv mos1S003-bkg.pi mos1S003-bkg-180-240.pi
mv mos1S003.rmf mos1S003-180-240.rmf
mv mos1S003.arf mos1S003-180-240.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-180-240.fits
specgroup spectrumset=mos1S003-obj-180-240.pi mincounts=100 rmfset=mos1S003-180-240.rmf \
arfset=mos1S003-180-240.arf backgndset=mos1S003-bkg-180-240.pi \
groupedset=mos1S003-obj-180-240-grp.pimosspectra eventfile=mos2S004-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg2-180-240.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-180-240.pi
mv mos2S004-bkg.pi mos2S004-bkg-180-240.pi
mv mos2S004.rmf mos2S004-180-240.rmf
mv mos2S004.arf mos2S004-180-240.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-180-240.fits
specgroup spectrumset=mos2S004-obj-180-240.pi mincounts=100 rmfset=mos2S004-180-240.rmf \
arfset=mos2S004-180-240.arf backgndset=mos2S004-bkg-180-240.pi \
groupedset=mos2S004-obj-180-240-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-180-240.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-180-240.pi
mv pnS005-fovt.pi pnS005-obj-180-240.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-180-240.pi
mv pnS005-bkg.pi pnS005-bkg-180-240.pi
mv pnS005.rmf pnS005-180-240.rmf
mv pnS005.arf pnS005-180-240.arf
mv pnS005-fovimspdet.fits pnS005-sp-180-240.fits
specgroup spectrumset=pnS005-obj-180-240.pi mincounts=100 rmfset=pnS005-180-240.rmf \
arfset=pnS005-180-240.arf backgndset=pnS005-bkg-180-240.pi \
groupedset=pnS005-obj-180-240-grp.pimosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-240-300.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-240-300.pi
mv mos1S003-bkg.pi mos1S003-bkg-240-300.pi
mv mos1S003.rmf mos1S003-240-300.rmf
mv mos1S003.arf mos1S003-240-300.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-240-300.fits
specgroup spectrumset=mos1S003-obj-240-300.pi mincounts=100 rmfset=mos1S003-240-300.rmf \
arfset=mos1S003-240-300.arf backgndset=mos1S003-bkg-240-300.pi \
groupedset=mos1S003-obj-240-300-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-240-300.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-240-300.pi
mv mos2S004-bkg.pi mos2S004-bkg-240-300.pi
mv mos2S004.rmf mos2S004-240-300.rmf
mv mos2S004.arf mos2S004-240-300.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-240-300.fits
specgroup spectrumset=mos2S004-obj-240-300.pi mincounts=100 rmfset=mos2S004-240-300.rmf \
arfset=mos2S004-240-300.arf backgndset=mos2S004-bkg-240-300.pi \
groupedset=mos2S004-obj-240-300-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-240-300.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-240-300.pi
mv pnS005-fovt.pi pnS005-obj-240-300.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-240-300.pi
mv pnS005-bkg.pi pnS005-bkg-240-300.pi
mv pnS005.rmf pnS005-240-300.rmf
mv pnS005.arf pnS005-240-300.arf
mv pnS005-fovimspdet.fits pnS005-sp-240-300.fits
specgroup spectrumset=pnS005-obj-240-300.pi mincounts=100 rmfset=pnS005-240-300.rmf \
arfset=pnS005-240-300.arf backgndset=pnS005-bkg-240-300.pi \
groupedset=pnS005-obj-240-300-grp.piSkip to the end of the this section.
mosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-300-420.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-300-420.pi
mv mos1S003-bkg.pi mos1S003-bkg-300-420.pi
mv mos1S003.rmf mos1S003-300-420.rmf
mv mos1S003.arf mos1S003-300-420.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-300-420.fits
specgroup spectrumset=mos1S003-obj-300-420.pi mincounts=100 rmfset=mos1S003-300-420.rmf \
arfset=mos1S003-300-420.arf backgndset=mos1S003-bkg-300-420.pi \
groupedset=mos1S003-obj-300-420-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-300-420.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-300-420.pi
mv mos2S004-bkg.pi mos2S004-bkg-300-420.pi
mv mos2S004.rmf mos2S004-300-420.rmf
mv mos2S004.arf mos2S004-300-420.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-300-420.fits
specgroup spectrumset=mos2S004-obj-300-420.pi mincounts=100 rmfset=mos2S004-300-420.rmf \
arfset=mos2S004-300-420.arf backgndset=mos2S004-bkg-300-420.pi \
groupedset=mos2S004-obj-300-420-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-300-420.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-300-420.pi
mv pnS005-fovt.pi pnS005-obj-300-420.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-300-420.pi
mv pnS005-bkg.pi pnS005-bkg-300-420.pi
mv pnS005.rmf pnS005-300-420.rmf
mv pnS005.arf pnS005-300-420.arf
mv pnS005-fovimspdet.fits pnS005-sp-300-420.fits
specgroup spectrumset=pnS005-obj-300-420.pi mincounts=100 rmfset=pnS005-300-420.rmf \
arfset=pnS005-300-420.arf backgndset=pnS005-bkg-300-420.pi \
groupedset=pnS005-obj-300-420-grp.pimosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-420-540.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi elow=300 ehigh=1100 ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-420-540.pi
mv mos1S003-bkg.pi mos1S003-bkg-420-540.pi
mv mos1S003.rmf mos1S003-420-540.rmf
mv mos1S003.arf mos1S003-420-540.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-420-540.fits
specgroup spectrumset=mos1S003-obj-420-540.pi mincounts=100 rmfset=mos1S003-420-540.rmf \
arfset=mos1S003-420-540.arf backgndset=mos1S003-bkg-420-540.pi \
groupedset=mos1S003-obj-420-540-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-420-540.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-420-540.pi
mv mos2S004-bkg.pi mos2S004-bkg-420-540.pi
mv mos2S004.rmf mos2S004-420-540.rmf
mv mos2S004.arf mos2S004-420-540.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-420-540.fits
specgroup spectrumset=mos2S004-obj-420-540.pi mincounts=100 rmfset=mos2S004-420-540.rmf \
arfset=mos2S004-420-540.arf backgndset=mos2S004-bkg-420-540.pi \
groupedset=mos2S004-obj-420-540-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-420-540.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-420-540.pi
mv pnS005-fovt.pi pnS005-obj-420-540.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-420-540.pi
mv pnS005-bkg.pi pnS005-bkg-420-540.pi
mv pnS005.rmf pnS005-420-540.rmf
mv pnS005.arf pnS005-420-540.arf
mv pnS005-fovimspdet.fits pnS005-sp-420-540.fits
specgroup spectrumset=pnS005-obj-420-540.pi mincounts=100 rmfset=pnS005-420-540.rmf \
arfset=pnS005-420-540.arf backgndset=pnS005-bkg-420-540.pi \
groupedset=pnS005-obj-420-540-grp.pimosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-540-720.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile
mv mos1S003-fovt.pi mos1S003-obj-540-720.pi
mv mos1S003-bkg.pi mos1S003-bkg-540-720.pi
mv mos1S003.rmf mos1S003-540-720.rmf
mv mos1S003.arf mos1S003-540-720.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-540-720.fits
specgroup spectrumset=mos1S003-obj-540-720.pi mincounts=100 rmfset=mos1S003-540-720.rmf \
arfset=mos1S003-540-720.arf backgndset=mos1S003-bkg-540-720.pi \
groupedset=mos1S003-obj-540-720-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-540-720.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-540-720.pi
mv mos2S004-bkg.pi mos2S004-bkg-540-720.pi
mv mos2S004.rmf mos2S004-540-720.rmf
mv mos2S004.arf mos2S004-540-720.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-540-720.fits
specgroup spectrumset=mos2S004-obj-540-720.pi mincounts=100 rmfset=mos2S004-540-720.rmf \
arfset=mos2S004-540-720.arf backgndset=mos2S004-bkg-540-720.pi \
groupedset=mos2S004-obj-540-720-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-540-720.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-540-720.pi
mv pnS005-fovt.pi pnS005-obj-540-720.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-540-720.pi
mv pnS005-bkg.pi pnS005-bkg-540-720.pi
mv pnS005.rmf pnS005-540-720.rmf
mv pnS005.arf pnS005-540-720.arf
mv pnS005-fovimspdet.fits pnS005-sp-540-720.fits
specgroup spectrumset=pnS005-obj-540-270.pi mincounts=100 rmfset=pnS005-540-720.rmf \
arfset=pnS005-540-720.arf backgndset=pnS005-bkg-540-720.pi \
groupedset=pnS005-obj-540-720-grp.pimosspectra eventfile=mos1S003-allevc.fits withregion=yes regionfile=reg1-720-840.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos1S003-fovt.pi ccds="T T T T T T T" \
mv mos1S003-fovt.pi mos1S003-obj-720-840.pi
mv mos1S003-bkg.pi mos1S003-bkg-720-840.pi
mv mos1S003.rmf mos1S003-720-840.rmf
mv mos1S003.arf mos1S003-720-840.arf
mv mos1S003-fovimspdet.fits mos1S003-sp-720-840.fits
specgroup spectrumset=mos1S003-obj-720-840.pi mincounts=100 rmfset=mos1S003-720-840.rmf \
arfset=mos1S003-720-840.arf backgndset=mos1S003-bkg-720-840.pi \
groupedset=mos1S003-obj-720-840-grp.pimosspectra eventfile=mos2S004-allevc.fits withregion=yes regionfile=reg2-720-840.txt \
pattern=12 ccds="T T T T T T T" withsrcrem=yes maskdet=-bkgregtdet.fits \
masksky=-bkgregtsky.fits
mosback inspecfile=mos2S004-fovt.pi ccds="T T T T T T T" \
mv mos2S004-fovt.pi mos2S004-obj-720-840.pi
mv mos2S004-bkg.pi mos2S004-bkg-720-840.pi
mv mos2S004.rmf mos2S004-720-840.rmf
mv mos2S004.arf mos2S004-720-840.arf
mv mos2S004-fovimspdet.fits mos2S004-sp-720-840.fits
specgroup spectrumset=mos2S004-obj-720-840.pi mincounts=100 rmfset=mos2S004-720-840.rmf \
arfset=mos2S004-720-840.arf backgndset=mos2S004-bkg-720-840.pi \
groupedset=mos2S004-obj-720-840-grp.pipnspectra eventfile=pnS005-allevc.fits ootevtfile=pnS005-allevcoot.fits \
withregion=yes regionfile=reg3-720-840.txt \
pattern=4 withsrcrem=yes maskdet=pnS005-bkgregtdet.fits masksky=pnS005-bkgregtsky.fits \
quads="T T T T"
pnback inspecfile=pnS005-fovt.pi inspecoot=pnS005-fovtoot.pi quads="T T T T" \
mv pnS005-obj-os.pi pnS005-obj-os-720-840.pi
mv pnS005-fovt.pi pnS005-obj-720-840.pi
mv pnS005-fovtoot.pi pnS005-obj-oot-720-840.pi
mv pnS005-bkg.pi pnS005-bkg-720-840.pi
mv pnS005.rmf pnS005-720-840.rmf
mv pnS005.arf pnS005-720-840.arf
mv pnS005-fovimspdet.fits pnS005-sp-720-840.fits
specgroup spectrumset=pnS005-obj-720-840.pi mincounts=100 rmfset=pnS005-720-840.rmf \
arfset=pnS005-720-840.arf backgndset=pnS005-bkg-720-840.pi \
groupedset=pnS005-obj-720-840-grp.pi
-
Because of XMM's finite PSF, if there are strong spectral gradients the spectrum in one region can be significantly affected by the spectrum in a neighboring region. The creation and use of cross arfs provides a means of addressing this cross-talk. Use arfgen and evselect to create cross arfs for the spectral analysis. evselect is first run to create an instrument map. Some care must be taken in choosing the pixel size of the image as arfgen computation time scale as the number of pixels but if the pixels are too coarse the ARF will be underestimated. Generally, the ARF will reach its asymptotic limit about the time it reached infinite computation time. After the detector map has been created, arfgen can be run for the appropriate ARFs - the smaller the region is, the finer the detector map should be. Also, not that arfgen in the cross-arf mode can be run for the appropriate ARF for a given region on its own.
Skip to the end of the this section.
evselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=50 ycolumn=DETY yimagebinsize=50 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-0-30-0-30.arf spectrumset=mos1S003-obj-0-30.pi \
crossreg_spectrumset=mos1S003-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-0-30-30-60.arf spectrumset=mos1S003-obj-30-60.pi \
crossreg_spectrumset=mos1S003-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-0-30-60-120.arf spectrumset=mos1S003-obj-60-120.pi \
crossreg_spectrumset=mos1S003-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-30-60-30-60.arf spectrumset=mos1S003-obj-30-60.pi \
crossreg_spectrumset=mos1S003-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-30-60-0-30.arf spectrumset=mos1S003-obj-0-30.pi \
crossreg_spectrumset=mos1S003-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-30-60-60-120.arf spectrumset=mos1S003-obj-60-120.pi \
crossreg_spectrumset=mos1S003-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=50 ycolumn=DETY yimagebinsize=50 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-0-30-0-30.arf spectrumset=mos2S004-obj-0-30.pi \
crossreg_spectrumset=mos2S004-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-0-30-30-60.arf spectrumset=mos2S004-obj-30-60.pi \
crossreg_spectrumset=mos2S004-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-0-30-60-120.arf spectrumset=mos2S004-obj-60-120.pi \
crossreg_spectrumset=mos2S004-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-30-60-30-60.arf spectrumset=mos2S004-obj-30-60.pi \
crossreg_spectrumset=mos2S004-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-30-60-0-30.arf spectrumset=mos2S004-obj-0-30.pi \
crossreg_spectrumset=mos2S004-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-30-60-60-120.arf spectrumset=mos2S004-obj-60-120.pi \
crossreg_spectrumset=mos2S004-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=50 ycolumn=DETY yimagebinsize=50 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-0-30-0-30.arf spectrumset=pnS005-obj-0-30.pi \
crossreg_spectrumset=pnS005-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-0-30-30-60.arf spectrumset=pnS005-obj-30-60.pi \
crossreg_spectrumset=pnS005-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-0-30-60-120.arf spectrumset=pnS005-obj-60-120.pi \
crossreg_spectrumset=pnS005-obj-0-30.pi crossregionarf=yes withrmfset=yes r\
mfset=pnS005-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-30-60-30-60.arf spectrumset=pnS005-obj-30-60.pi \
crossreg_spectrumset=pnS005-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-30-60-0-30.arf spectrumset=pnS005-obj-0-30.pi \
crossreg_spectrumset=pnS005-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-30-60-60-120.arf spectrumset=pnS005-obj-60-120.pi \
crossreg_spectrumset=pnS005-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=noSkip to the end of the this section.
evselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
cxcolumn=DETX ximagebinsize=70 ycolumn=DETY yimagebinsize=70 squarepixels=yes \
cimagebinning=binSize withimageset=yesarfgen arfset=mos1S003-0-30-120-180.arf spectrumset=mos1S003-obj-120-180.pi \
ccrossreg_spectrumset=mos1S003-obj-0-30.pi crossregionarf=yes withrmfset=yes \
crmfset=mos1S003-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
cdetmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
cmodelootcorr=no
arfgen arfset=mos1S003-30-60-120-180.arf spectrumset=mos1S003-obj-120-180.pi \
ccrossreg_spectrumset=mos1S003-obj-30-60.pi crossregionarf=yes withrmfset=yes \
crmfset=mos1S003-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
cdetmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
cmodelootcorr=no
arfgen arfset=mos1S003-60-120-60-120.arf spectrumset=mos1S003-obj-60-120.pi \
ccrossreg_spectrumset=mos1S003-obj-60-120.pi crossregionarf=yes withrmfset=yes \
crmfset=mos1S003-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
cdetmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
cmodelootcorr=no
arfgen arfset=mos1S003-60-120-0-30.arf spectrumset=mos1S003-obj-0-30.pi \
ccrossreg_spectrumset=mos1S003-obj-60-120.pi crossregionarf=yes withrmfset=yes \
crmfset=mos1S003-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
cdetmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
cmodelootcorr=no
arfgen arfset=mos1S003-60-120-30-60.arf spectrumset=mos1S003-obj-30-60.pi \
crossreg_spectrumset=mos1S003-obj-60-120.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-60-120-120-180.arf spectrumset=mos1S003-obj-120-180.pi \
crossreg_spectrumset=mos1S003-obj-60-120.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=70 ycolumn=DETY yimagebinsize=70 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-0-30-120-180.arf spectrumset=mos2S004-obj-120-180.pi \
crossreg_spectrumset=mos2S004-obj-0-30.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-30-60-120-180.arf spectrumset=mos2S004-obj-120-180.pi \
crossreg_spectrumset=mos2S004-obj-30-60.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-60-120-60-120.arf spectrumset=mos2S004-obj-60-120.pi \
crossreg_spectrumset=mos2S004-obj-60-120.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-60-120-0-30.arf spectrumset=mos2S004-obj-0-30.pi \
crossreg_spectrumset=mos2S004-obj-60-120.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-60-120-30-60.arf spectrumset=mos2S004-obj-30-60.pi \
crossreg_spectrumset=mos2S004-obj-60-120.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-60-120-120-180.arf spectrumset=mos2S004-obj-120-180.pi \
crossreg_spectrumset=mos2S004-obj-60-120.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
mxcolumn=DETX ximagebinsize=70 ycolumn=DETY yimagebinsize=70 squarepixels=yes \
mimagebinning=binSize withimageset=yesarfgen arfset=pnS005-0-30-120-180.arf spectrumset=pnS005-obj-120-180.pi \
mcrossreg_spectrumset=pnS005-obj-0-30.pi crossregionarf=yes withrmfset=yes \
mrmfset=pnS005-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
mdetmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
mmodelootcorr=no
arfgen arfset=pnS005-30-60-120-180.arf spectrumset=pnS005-obj-120-180.pi \
mcrossreg_spectrumset=pnS005-obj-30-60.pi crossregionarf=yes withrmfset=yes \
mrmfset=pnS005-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
mdetmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
mmodelootcorr=no
arfgen arfset=pnS005-60-120-60-120.arf spectrumset=pnS005-obj-60-120.pi \
mcrossreg_spectrumset=pnS005-obj-60-120.pi crossregionarf=yes withrmfset=yes \
mrmfset=pnS005-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
mdetmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
mmodelootcorr=no
arfgen arfset=pnS005-60-120-0-30.arf spectrumset=pnS005-obj-0-30.pi \
mcrossreg_spectrumset=pnS005-obj-60-120.pi crossregionarf=yes withrmfset=yes \
mrmfset=pnS005-0-30.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
mdetmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
mmodelootcorr=no
arfgen arfset=pnS005-60-120-30-60.arf spectrumset=pnS005-obj-30-60.pi \
mcrossreg_spectrumset=pnS005-obj-60-120.pi crossregionarf=yes withrmfset=yes \
mrmfset=pnS005-30-60.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
mdetmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
mmodelootcorr=no
arfgen arfset=pnS005-60-120-120-180.arf spectrumset=pnS005-obj-120-180.pi \
mcrossreg_spectrumset=pnS005-obj-60-120.pi crossregionarf=yes withrmfset=yes \
mrmfset=pnS005-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
mdetmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
mmodelootcorr=noSkip to the end of the this section.
evselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=100 ycolumn=DETY yimagebinsize=100 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-120-180-120-180.arf spectrumset=mos1S003-obj-120-180.pi \
crossreg_spectrumset=mos1S003-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-120-180-60-120.arf spectrumset=mos1S003-obj-60-120.pi \
crossreg_spectrumset=mos1S003-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-120-180-180-240.arf spectrumset=mos1S003-obj-180-240.pi \
crossreg_spectrumset=mos1S003-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=100 ycolumn=DETY yimagebinsize=100 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-120-180-120-180.arf spectrumset=mos2S004-obj-120-180.pi \
crossreg_spectrumset=mos2S004-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-120-180-60-120.arf spectrumset=mos2S004-obj-60-120.pi \
crossreg_spectrumset=mos2S004-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-120-180-180-240.arf spectrumset=mos2S004-obj-180-240.pi \
crossreg_spectrumset=mos2S004-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=100 ycolumn=DETY yimagebinsize=100 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-120-180-120-180.arf spectrumset=pnS005-obj-120-180.pi \
crossreg_spectrumset=pnS005-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-120-180-60-120.arf spectrumset=pnS005-obj-60-120.pi \
crossreg_spectrumset=pnS005-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-60-120.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-120-180-180-240.arf spectrumset=pnS005-obj-180-240.pi \
crossreg_spectrumset=pnS005-obj-120-180.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=noevselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=150 ycolumn=DETY yimagebinsize=150 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-180-240-180-240.arf spectrumset=mos1S003-obj-180-240.pi \
crossreg_spectrumset=mos1S003-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-180-240-120-180.arf spectrumset=mos1S003-obj-120-180.pi \
crossreg_spectrumset=mos1S003-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-180-240-240-300.arf spectrumset=mos1S003-obj-240-300.pi \
crossreg_spectrumset=mos1S003-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=150 ycolumn=DETY yimagebinsize=150 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-180-240-180-240.arf spectrumset=mos2S004-obj-180-240.pi \
crossreg_spectrumset=mos2S004-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-180-240-120-180.arf spectrumset=mos2S004-obj-120-180.pi \
crossreg_spectrumset=mos2S004-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-180-240-240-300.arf spectrumset=mos2S004-obj-240-300.pi \
crossreg_spectrumset=mos2S004-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=150 ycolumn=DETY yimagebinsize=150 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-180-240-180-240.arf spectrumset=pnS005-obj-180-240.pi \
crossreg_spectrumset=pnS005-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-180-240-120-180.arf spectrumset=pnS005-obj-120-180.pi \
crossreg_spectrumset=pnS005-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-120-180.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-180-240-240-300.arf spectrumset=pnS005-obj-240-300.pi \
crossreg_spectrumset=pnS005-obj-180-240.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=noSkip to the end of the this section.
evselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=200 ycolumn=DETY yimagebinsize=200 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-240-300-240-300.arf spectrumset=mos1S003-obj-240-300.pi \
crossreg_spectrumset=mos1S003-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-240-300-180-240.arf spectrumset=mos1S003-obj-180-240.pi \
crossreg_spectrumset=mos1S003-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-240-300-300-420.arf spectrumset=mos1S003-obj-300-420.pi \
crossreg_spectrumset=mos1S003-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=200 ycolumn=DETY yimagebinsize=200 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-240-300-240-300.arf spectrumset=mos2S004-obj-240-300.pi \
crossreg_spectrumset=mos2S004-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-240-300-180-240.arf spectrumset=mos2S004-obj-180-240.pi \
crossreg_spectrumset=mos2S004-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-240-300-300-420.arf spectrumset=mos2S004-obj-300-420.pi \
crossreg_spectrumset=mos2S004-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=200 ycolumn=DETY yimagebinsize=200 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-240-300-240-300.arf spectrumset=pnS005-obj-240-300.pi \
crossreg_spectrumset=pnS005-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-240-300-180-240.arf spectrumset=pnS005-obj-180-240.pi \
crossreg_spectrumset=pnS005-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-180-240.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-240-300-300-420.arf spectrumset=pnS005-obj-300-420.pi \
crossreg_spectrumset=pnS005-obj-240-300.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=noevselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=250 ycolumn=DETY yimagebinsize=250 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-300-420-300-420.arf spectrumset=mos1S003-obj-300-420.pi \
crossreg_spectrumset=mos1S003-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-300-420-240-300.arf spectrumset=mos1S003-obj-240-300.pi \
crossreg_spectrumset=mos1S003-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-300-420-420-540.arf spectrumset=mos1S003-obj-420-540.pi \
crossreg_spectrumset=mos1S003-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=250 ycolumn=DETY yimagebinsize=250 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-300-420-300-420.arf spectrumset=mos2S004-obj-300-420.pi \
crossreg_spectrumset=mos2S004-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-300-420-240-300.arf spectrumset=mos2S004-obj-240-300.pi \
crossreg_spectrumset=mos2S004-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-300-420-420-540.arf spectrumset=mos2S004-obj-420-540.pi \
crossreg_spectrumset=mos2S004-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=250 ycolumn=DETY yimagebinsize=250 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-300-420-300-420.arf spectrumset=pnS005-obj-300-420.pi \
crossreg_spectrumset=pnS005-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-300-420-240-300.arf spectrumset=pnS005-obj-240-300.pi \
crossreg_spectrumset=pnS005-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-240-300.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-300-420-420-540.arf spectrumset=pnS005-obj-420-540.pi \
crossreg_spectrumset=pnS005-obj-300-420.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=noSkip to the end of the this section.
evselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=300 ycolumn=DETY yimagebinsize=300 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-420-540-420-540.arf spectrumset=mos1S003-obj-420-540.pi \
crossreg_spectrumset=mos1S003-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits\
modelootcorr=no
arfgen arfset=mos1S003-420-540-300-420.arf spectrumset=mos1S003-obj-300-420.pi \
crossreg_spectrumset=mos1S003-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-420-540-540-720.arf spectrumset=mos1S003-obj-540-720.pi \
crossreg_spectrumset=mos1S003-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=300 ycolumn=DETY yimagebinsize=300 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-420-540-420-540.arf spectrumset=mos2S004-obj-420-540.pi \
crossreg_spectrumset=mos2S004-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-420-540-300-420.arf spectrumset=mos2S004-obj-300-420.pi \
crossreg_spectrumset=mos2S004-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-420-540-540-720.arf spectrumset=mos2S004-obj-540-720.pi \
crossreg_spectrumset=mos2S004-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=300 ycolumn=DETY yimagebinsize=300 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-420-540-420-540.arf spectrumset=pnS005-obj-420-540.pi \
crossreg_spectrumset=pnS005-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-420-540-300-420.arf spectrumset=pnS005-obj-300-420.pi \
crossreg_spectrumset=pnS005-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-300-420.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-420-540-540-720.arf spectrumset=pnS005-obj-540-720.pi \
crossreg_spectrumset=pnS005-obj-420-540.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=noevselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=350 ycolumn=DETY yimagebinsize=350 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-540-720-540-720.arf spectrumset=mos1S003-obj-540-720.pi \
crossreg_spectrumset=mos1S003-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-540-720-420-540.arf spectrumset=mos1S003-obj-420-540.pi \
crossreg_spectrumset=mos1S003-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-540-720-720-840.arf spectrumset=mos1S003-obj-720-840.pi \
crossreg_spectrumset=mos1S003-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-720-840.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=350 ycolumn=DETY yimagebinsize=350 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-540-720-540-720.arf spectrumset=mos2S004-obj-540-720.pi \
crossreg_spectrumset=mos2S004-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-540-720-420-540.arf spectrumset=mos2S004-obj-420-540.pi \
crossreg_spectrumset=mos2S004-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-540-720-720-840.arf spectrumset=mos2S004-obj-720-840.pi \
crossreg_spectrumset=mos2S004-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-720-840.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=350 ycolumn=DETY yimagebinsize=350 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-540-720-540-720.arf spectrumset=pnS005-obj-540-720.pi \
crossreg_spectrumset=pnS005-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-540-720-420-540.arf spectrumset=pnS005-obj-420-540.pi \
crossreg_spectrumset=pnS005-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-420-540.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-540-720-720-840.arf spectrumset=pnS005-obj-720-840.pi \
crossreg_spectrumset=pnS005-obj-540-720.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-720-840.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=noSkip to the end of the this section.
evselect table=mos1S003-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=400 ycolumn=DETY yimagebinsize=400 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos1S003-720-840-720-840.arf spectrumset=mos1S003-obj-720-840.pi \
crossreg_spectrumset=mos1S003-obj-720-840.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-720-840.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=no
arfgen arfset=mos1S003-720-840-540-720.arf spectrumset=mos1S003-obj-540-720.pi \
crossreg_spectrumset=mos1S003-obj-720-840.pi crossregionarf=yes withrmfset=yes \
rmfset=mos1S003-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos1S003-clean.fits \
modelootcorr=noevselect table=mos2S004-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=400 ycolumn=DETY yimagebinsize=400 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=mos2S004-720-840-720-840.arf spectrumset=mos2S004-obj-720-840.pi \
crossreg_spectrumset=mos2S004-obj-720-840.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-720-840.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=no
arfgen arfset=mos2S004-720-840-540-720.arf spectrumset=mos2S004-obj-540-720.pi \
crossreg_spectrumset=mos2S004-obj-720-840.pi crossregionarf=yes withrmfset=yes \
rmfset=mos2S004-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=mos2S004-clean.fits \
modelootcorr=noevselect table=pnS005-clean.fits:EVENTS ignorelegallimits=yes imageset=detmap.ds \
xcolumn=DETX ximagebinsize=400 ycolumn=DETY yimagebinsize=400 squarepixels=yes \
imagebinning=binSize withimageset=yesarfgen arfset=pnS005-720-840-720-840.arf spectrumset=pnS005-obj-720-840.pi \
crossreg_spectrumset=pnS005-obj-720-840.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-720-840.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
arfgen arfset=pnS005-720-840-540-720.arf spectrumset=pnS005-obj-540-720.pi \
crossreg_spectrumset=pnS005-obj-720-840.pi crossregionarf=yes withrmfset=yes \
rmfset=pnS005-540-720.rmf extendedsource=yes modelee=no withbadpixcorr=yes \
detmaptype=dataset detmaparray=detmap.ds badpixlocation=pnS005-clean.fits \
modelootcorr=no
-
At this point all of the XMM spectra (source and background) and calibration files (RMF and ARF) files have been created for the spectral fitting process. However, it must be remembered that there are other background components besides the QPB which still contribute to the source spectrum: the cosmic X-ray background and possibly solar wind charge exchange (typically only the OVII and OVIII lines are critical) and soft proton events. Each of these three background components can be addressed as part of the fitting process. For example (in pseudo Xspec terms):
Model = cosmic + swcx + sp + source
= (apec + wabs*(apec + apec + pow) + (gauss + gauss) + (pow) + (source)
-> conUnfortunately, the fitting is more complicated than this. In Xspec V12.0 and higher, the soft proton component must be added as a separate model as it requires a separate diagonal response file. The cosmic background component needs to be constrained, which can be accomplished by simultaneously fitting a ROSAT All-Sky Survey (RASS) spectrum of the off-source background. The RASS spectrum and associated response file can be obtained using the HEASARC X-ray Background Tool. As extended emission is being studied (or you wouldn't be looking at this script), the various spectra need to be normalized to the same solid angle, per square arc minute, which is useful as the RASS spectra come in those units. A second constant scale factor can be added to the model and allowed to float to account for residual calibration discrepancies. Finally, additional Gaussian lines must be added to the model to account for the instrumental fluorescent lines associated with the QPD but are not in the model QPB spectra (the lines are so bright that the residual gain variation of an observation will cause significant residuals).
Model = cosmic + swcx + sp + source
= (apec + wabs*(apec + apec + pow) + (gauss + gauss) + (pow) + (source)
-> (gauss + gauss + gauss + gauss + gauss + gauss + gauss) +
con*con(gauss + gauss + apec + wabs*(apec + apec + pow) + source)To further complicate the model setup, all the spectra should be fit simultaneously to maximize the available constraints, e.g., the cosmic and SWCX background are the same for all XMM-Newton spectra and some parameters of the fluorescent line and SP background can be linked. This is all discussed to a much greater extent in the XMM-ESAS Cookbook.
A sample Xspec XCM file for these data can be found here. Figure 5 shows the results of the fit.
|
For information about known issues, refer to the ESAS Warnings and Watchouts pages. |
- Removed a total of (1) style text-align:center;
- Removed a total of (43) style text-align:justify;
- Removed a total of (1) style margin:0;
- Removed a total of (2) border attribute.
- Converted a total of (9) center to div.






































 Sign in
Sign in
 Science & Technology
Science & Technology