SAS Thread - epic merging - XMM-Newton
Combining the spectra of the 3 EPIC cameras
|
Introduction This thread is a step-by-step recipe to combine the spectra of a given number of EPIC camera exposures into one single spectrum with corresponding rmf, arf and bkg files. Expected Outcome The thread requires as input a set of spectral files from a given number of EPIC exposures. In this thread the source, background, response and effective area files are called: src_spectrum_#.ds, bkg_spectrum_#.ds, response_#.rmf and arf_#.arf, where # is used to indicate the spectrum set corresponding to a given EPIC exposure. It is assumed that the sets of spectral files have been produced prior to starting this thread. This thread shows how spectra and response files are merged together into one single set. The output of this thread is a merged source+background spectrum, background spectrum, redistribution matrix and effective area files. SAS Tasks to be Used Prerequisites
Useful Links Last Reviewed: 31 January 2025, for SAS v22.0Last Updated: 03 December 2021 |
Procedure
This thread assumes as starting point one set of spectral files for each one of the EPIC exposures that are going to be merged. For each EPIC exposure, a spectral set is made out of source+background spectrum, background spectrum, response and effective area files.
We note that since SAS v20.0 it is no longer necessary to extract all spectra over the same channel / energy interval. In fact, we recommend strongly against doing this, as this will lead to a wrong energy scale at low energies. The only requirment is that the RMF is being extracted with the same bin width, as outlined below. The thread below reflects the new and correct way of extracting and combining spectra.
Please find below a detailed walk-through:
Generation of individual exposure spectra and responses
The following three steps have to be repeated for every single spectral set to be generated.
- For each EPIC exposure generate a source+background spectrum in the standard PI range, that is (0-20479) for EPIC-pn and (0-11999) for EPIC-MOS. Use a common bin size of 5 eV.
evselect table=Events_01.evt expression="${src_exp}" \
withspectrumset=yes spectrumset=src_spectrum_01.ds \
spectralbinsize=5 specchannelmin=0 specchannelmax=${MAXCHAN} energycolumn=PIbackscale spectrumset=src_spectrum_01.ds badpixlocation=Events_01.evt
where ${src_exp} is a selectlib expression appropriate for the camera and source of interest and ${MAXCHAN} is 20479 for pn and 11999 for MOS. The threads on MOS [pn] show how to define expressions for spectral extraction with SAS.
- For each EPIC exposure generate a background spectrum with the same spectral range and with the same spectral binning as the one used in Step#1 above.
evselect table=Events_01.evt expression="${bkg_exp}" \
withspectrumset=yes spectrumset=bkg_spectrum_01.ds \
spectralbinsize=5 specchannelmin=0 specchannelmax=${MAXCHAN} energycolumn=PIbackscale spectrumset=bkg_spectrum_01.ds badpixlocation=Events_01.evt
where ${bkg_exp} is the selectlib expression appropriate to select the background for the source of interest.
- Generate redistribution matrices corresponding to the spectra extracted in Step#1.
rmfgen spectrumset=src_spectrum_01.ds rmfset=response_01.rmf \
withenergybins=yes energymin=0.1 energymax=${MAXENERGY} nenergybins=${NBINS}where ${MAXENERGY} is 15 for EPIC-pn and 12 for EPIC-MOS and correspondingly ${NBINS} is 1490 for pn and 1190 for MOS. It is important to adapt ${MAXENERGY} and ${NBINS} in such a way that they result in the same bin width for all spectra (0.01 in this case). It is also possible to cutoff the RMF generation for pn at 12keV (${MAXENERGY}=12 ), in that case also you need to also set $NBINS=1190 and acceptchanrange=yes.
- Generate the corresponding effective area files.
arfgen spectrumset=src_spectrum_01.ds arfset=arf_01.arf withrmfset=yes rmfset=response_01.rmf
In Step#1-Step#4 a set of spectral files has been generated for a given EPIC exposure, 01 in this example: src_spectrum_01.ds, bkg_spectrum_01.ds, response_01.rmf and arf_01.arf. Follow the same process to generate a spectral set for the rest of the EPIC exposures to be combined.
Combine the EPIC camera exposures spectra
For the purpose of this thread, lets assume that we have three EPIC exposures (01, 02, 03). This can be in any combination of pn or MOS camera exposures from the same or from different ODFs. Note that the tool requires the exposures to be ordered by instrument, e.g., all MOS1 spectra to be combined have to appear consecutively, the same is true for MOS2 and PN, independently of whether they belong or not to the same ODFs.
The call to epicspeccombine would look like this,
epicspeccombine pha="src_spectrum_01.ds src_spectrum_02.ds src_spectrum_03.ds"\
bkg="bkg_spectrum_01.ds bkg_spectrum_02.ds bkg_spectrum_03.ds" \
rmf="response_01.rmf response_02.rmf response_03.rmf" \
arf="arf_01.arf arf_02.arf arf_03.arf" \
filepha="src_spectrum_grp.ds" \
filebkg="bkg_spectrum_grp.ds" \
filersp="response_grp.rmf" \
allowHEdiff=yes
Where the three output files, src_spectrum_grp.ds, bkg_spectrum_grp.ds and response_grp.rmf correspond to the merged spectrum. The input response matrix and effective area files are now combined in a single response matrix, response_grp.rmf, hence, no arf file is generated for the merged spectrum. These files are ready to be loaded in packages like xspec.
The allowHEdiff=yes keyword is important to allow the combination of different energy ranges of the input spectra, e.g., to combine pn and MOS spectra extracted as described above.
Further grouping of the spectral files is possible with, for example, the SAS task specgroup . In this case a spectrum produced by the combination of the EPIC cameras will be grouped assuming that the detector response function is that of MOS-1.
Last, fit the merged spectrum.
|
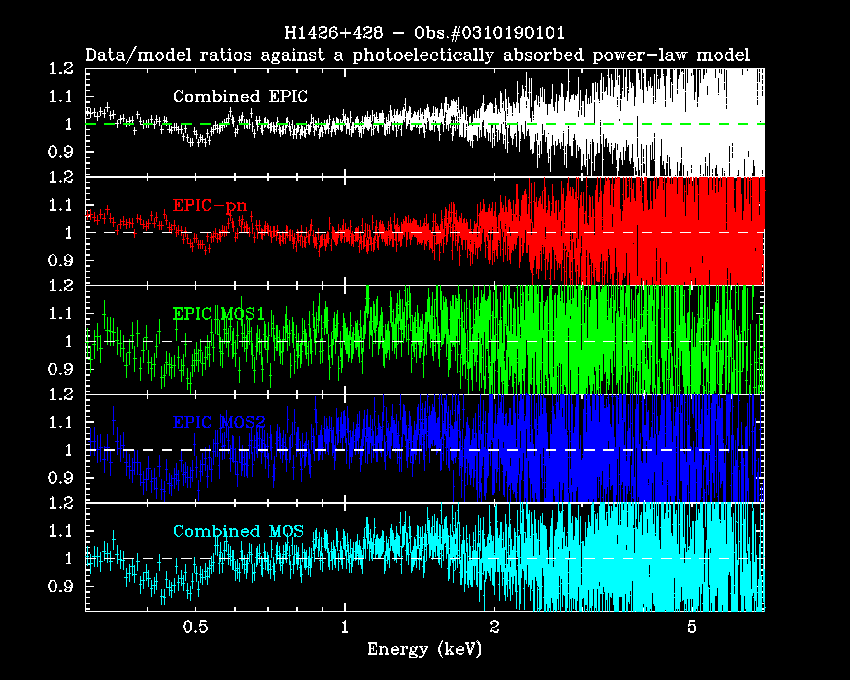
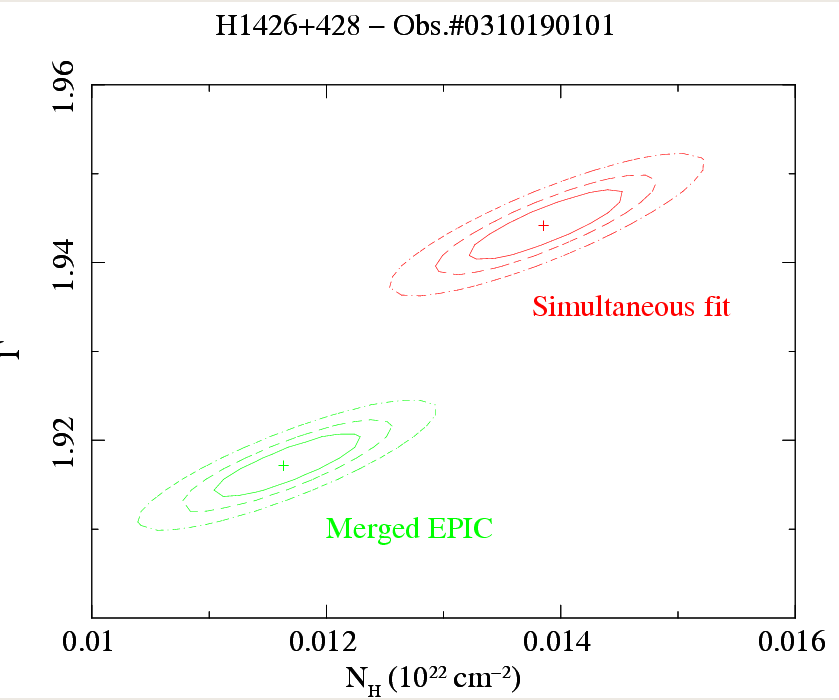
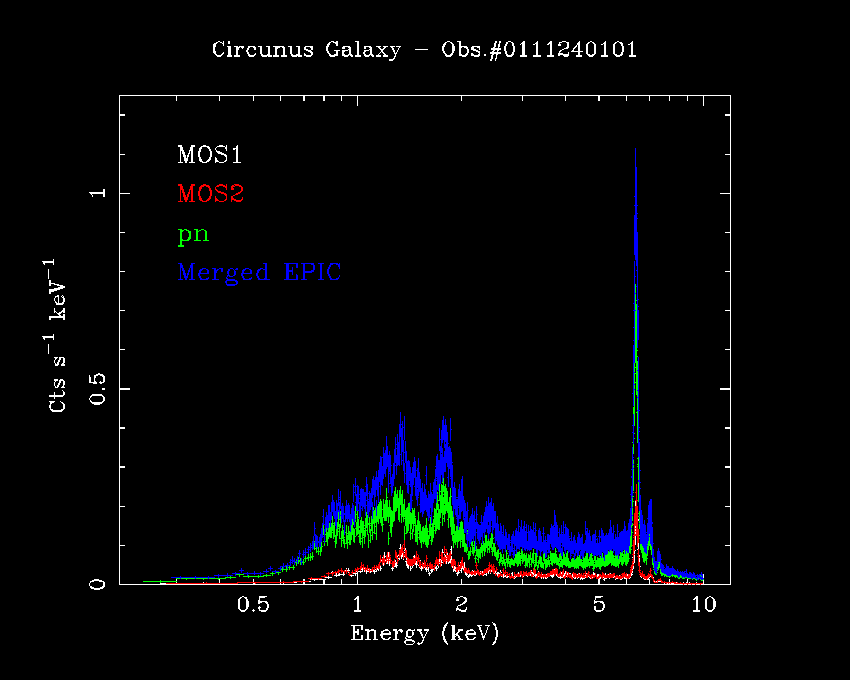
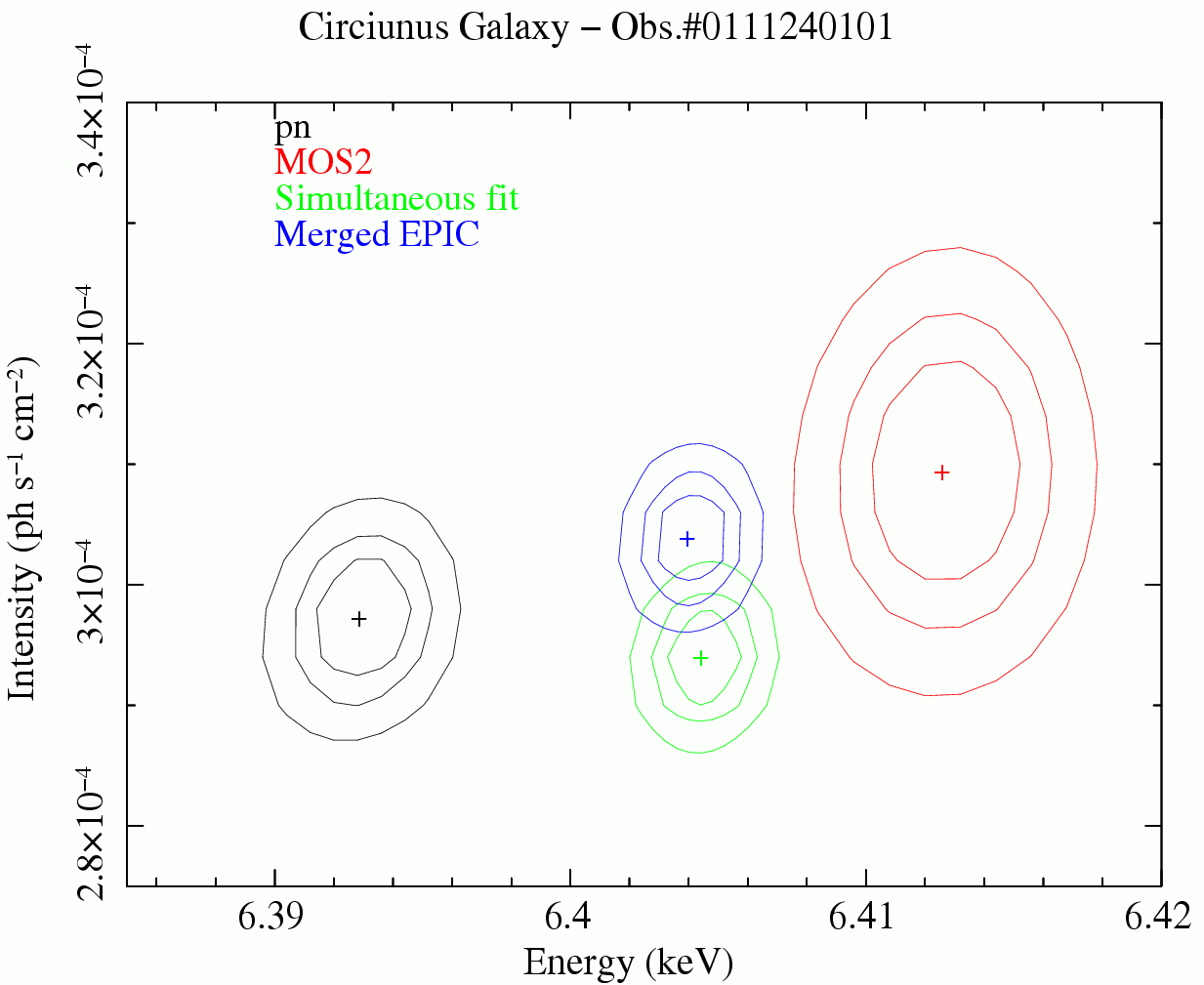
Users must be careful in using the algorithm and scripts described in this thread, above all, with bright sources whose spectral residuals are dominated by systematic errors. An example of the application of this script to real data (the EPIC spectra of the blazar H1426+428, Obs.#0310190101) is shown in Fig.2. The statistical quality of the parameters derived from the merged EPIC spectrum is better than the fit on one individual camera, but still comparable to that obtained by the simultaneous fitting of individual camera spectra, the latter being the rigorously correct procedure. Similar conclusions can be drawn with respect to the measurements of spectral properties of narrow-band features (in Fig.3 the prominent iron Kα fluorescent line in the EPIC spectra of the Circinus Galaxy, Molendi et al., 2003, MNRAS, 343, L1)
An alternative method to producing the merge spectrum files is the use of the task multiespecget. This tasks starts with the event files and will produce the files, src_spectrum_grp.ds, bkg_spectrum_grp.ds and response_grp.rmf corresponding to the merged spectrum. multiespecget should be run in this case with withmultiEPICspec=yes. IMPORTANT: the current implementation of multiespecget uses a different implementation than described here, in partciular it produces pn spectra with spechannelmax =11999. As this is a non-standard extraction range, rmfgen cannot produce a correct RMF and hence the energy-scale (in partiuclar at low energies) will be wrong. This behavior should be changed in an upcoming SAS release. |
- Removed a total of (4) style text-align:center;
- Removed a total of (16) style text-align:justify;
- Removed a total of (6) align=center.
- Removed a total of (2) border attribute.
- Converted a total of (2) center to div.






































 Sign in
Sign in
 Science & Technology
Science & Technology